Past Projects (III)
Rotaplotype was a java program asked by my former thesis master Dr D Poncet in order to find and visalize compensatory co-mutations in multiple rotavirus aligned genomes. The program called Rotaplotype took as input a set of rotavirus sequence aligned with clustalw and optionaly a RNAML-based description of the secondary structure of one viral RNA fragment.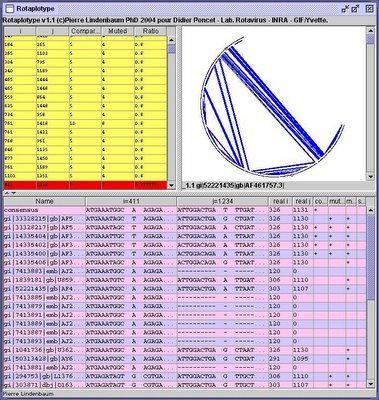
- on the top-left pane is displayed each co-mutation found in the alignments.
- on the top-right pane is displayed the position of the two mutations in the structure (may be of interest if the two mutations are localized in the same hairpin)
- on bottom are displayed the context of the two mutation for each sequence aligned by clustalw.
... not sure Dr Poncet discovered the deep secrets of the universe with this small tool
;-).
No comments:
Post a Comment