If you're following me on twitter or on friendfeed you may know that I've re-written a new version of SciFOAF.
Here is the documentation:
What is SciFOAF
SciFOAF is the second version of a tool I created to build a
FOAF/RDF file from your publications in
ncbi/pubmed. The FOAF project defines a semantic format based on RDF/XML to define persons or groups, their relationships, as well as their basic properties such as name, e-mail address, subjects of interest, publications, and so on... This FOAF profile can be used to describe your work, your laboratory, your contacts.
The first version was introduced in 2006
here as a java webstart interface and had many problems:
- the RDF file could not be loaded/saved
- only a few properties could be edited
- authors'name definition may vary from one journal to another as some journal may use the initial of an author while another may use the complete first name.
- the interaction was just a kind of multiple-choice questionnaire
The new version now uses the
Jena API, the rdf repository can be loaded and saved.
Requirements
Downloading SciFOAF
A *.jar file should be available for download at
http://lindenb.googlecode.com/files/scifoaf.jar.
Running SciFOAF
Setup the CLASSPATH
export JENA_LIB=your_path_to/Jena/lib
export CLASSPATH=${JENA_LIB}/antlr-2.7.5.jar:${JENA_LIB}/arq-extra.jar:${JENA_LIB}/arq.jar:${JENA_LIB}/commons-logging-1.1.1.jar:${JENA_LIB}/concurrent.jar:${JENA_LIB}/icu4j_3_4.jar:${JENA_LIB}/iri.jar:${JENA_LIB}/jena.jar:${JENA_LIB}/jenatest.jar:${JENA_LIB}/json.jar:${JENA_LIB}/junit.jar:${JENA_LIB}/log4j-1.2.12.jar:${JENA_LIB}/lucene-core-2.3.1.jar:${JENA_LIB}/stax-api-1.0.jar:${JENA_LIB}/wstx-asl-3.0.0.jar:${JENA_LIB}/xercesImpl.jar:${JENA_LIB}/xml-apis.jar:YOUR_PATH_TO/scifoaf.jar
Run SciFOAF
java org.lindenb.scifoaf.SciFOAF
the first time your run SciFOAF, You're prompted to give yourself an URI. The best choice would be to give the URL where your foaf file will be stored or the URL of your personnal homepage or blog. On startup a file called
foaf.rdf will be created in your home directory. Alternatively you can specify a file on the command line.
When the application is closed, the FOAF model will be saved back to the file.
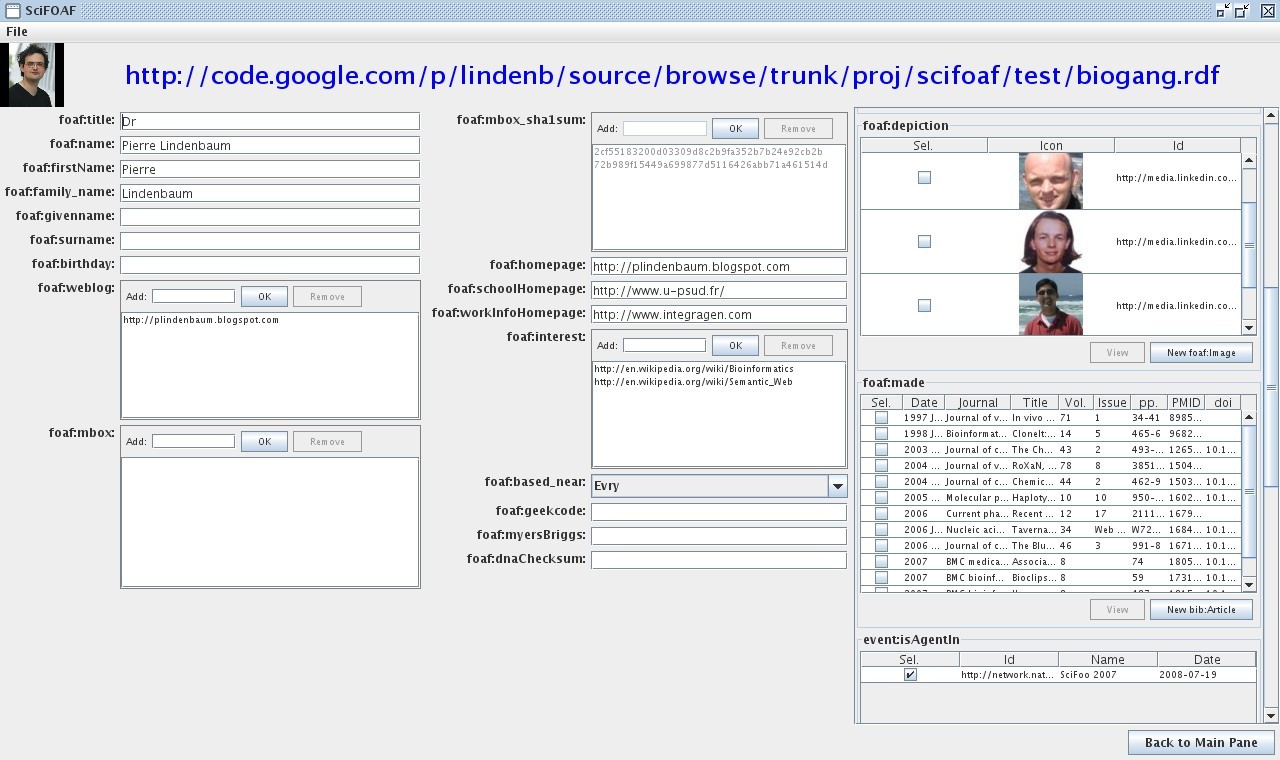
The Main Pane
The first window contains a sequence of tab Each tab fits to a given rdf Class:
- foaf:Person
- geo:Place
- bibo:Article
- ...
For each tab, a button "New ...." creates a new instance of the given Class.
Building your profile
Add a foaf:Image
Add the URL of the picture, for example:
http://upload.wikimedia.org/wikipedia/commons/4/42/Charles_Darwin_aged_51.jpg.
Add an bibo:Article
enter the PMID of the artcle
Add a geo:Place
SciFOAF, uses the
geonames.org API.
Add a foaf:Person
You can the link this person to his publication, his foaf:based_near, the persons he knows..
Etc...
Create foaf:Group, event:Event, doap:Project....
Exporting to KML
(Experimental) In menu "File' select 'Export to KML'. SciFOAF will export a
KML file containing the geolocalized foaf:Persons.
A test is available
here and is visible in maps.google.com at
http://maps.google.com/maps?q=http://yokofakun.....
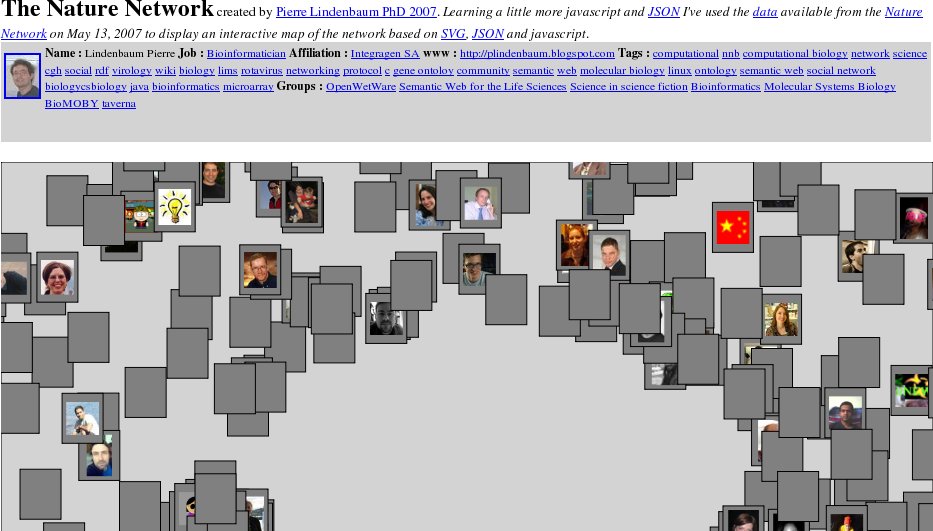
Exporting to XHTML+SVG
(Experimental) In menu "File' select 'Export to XHTML'. Here, I've roughly copied the tool I wrote for exploring the
Nature Network using SVG/javscript/JSON/XTML. Many things remain to do.
Loading a Batch of Articles
In the main panel, for
bibo:Article a button can be used to load a batch of articles.
On ncbi/pubmed, perform a query, choose
Example
A RDF File describing a few persons in the
Biogang is available
here.
Source Code
The source code is available on
http://code.google.com/p/lindenb/.
The ant file is in
lindenb/proj/scifoaf/build.xml
.
Pierre